Our analytical methods are modular and can be combined to great advantage.
Choose one of our services to learn more about the method.
Our goal is to answer your questions.
The polymerase chain reaction (PCR) is the most fundamental and important working method of IDENTXX.
Four components are needed for a successful PCR test
PCR procedure
For qualitative detection, also known as end-point PCR, the enzyme reaction is first carried out completely. After the reaction is complete, the resulting products are separated according to size in an agarose gel and visualized using fluorescent dyes.
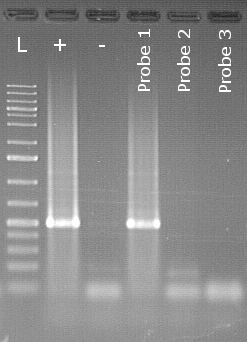
Specific endpoint PCR detection of Peronospora belbahrii
The example shows an agarose gel used to detect Peronospora belbahrii,
the downy mildew of basil, from seed. Next to the
size marker (L) is the positive control (+) followed by the negative control (-).
In the positive control lane, a clearly visible bright white area,
also known as a "band," can be seen. This is the stained specific PCR product. A band is also visible at the same level in the lane of sample 1. This sample is therefore considered positive. Samples 2 and 3 show no bands and are therefore considered negative.
Our qualitative PCR tests are safe and robust, and are used to detect a wide variety of pathogens.
Another important PCR method is quantitative PCR (qPCR), also known as real-time PCR. It follows the same reaction principles as qualitative PCR. However, the reaction solution is supplemented with an additional component, the so-called probe. This probe is located between the primers and is also specific for the DNA segment being sought.
When a PCR reaction takes place, the polymerase enzyme breaks down the probe. A fluorescent dye (fluorophore) released in this process can be detected in real time. The more PCR products are formed, the more fluorophore is released. The thermocycler displays these measurements as a curve. The sooner the curve shows an exponential rise, the more DNA was present in the initial solution.
The use of probes offers another advantage. By using different fluorophores, we can perform up to five PCR reactions in a single reaction vessel (multiplex analyses). This makes it possible to compare the concentration of specific pathogen DNA (shown in blue in the upper curve) to the DNA of the host plant (shown in green).
This approach has several advantages:
We also used endpoint PCR as a basis for DNA sequencing, distinguishing between Sanger sequencing and pyrosequencing. These are two important analytical tools that we used for different analytical purposes.
This method, developed by Frederick Sanger and Alan Coulson in the 1970s, forms the foundation of modern molecular biology alongside PCR. The method is as simple as it is ingenious. Special oligonucleotides labeled with fluorescent dyes are added to the prepared endpoint PCR reaction solution. In a subsequent PCR reaction, DNA fragments of varying lengths are obtained, each ending with a fluorescently labeled oligonucleotide that can be analyzed using appropriate detectors. The result is a sequence of colored tips corresponding to the four bases of DNA.
Pyrosequencing is based on endpoint PCR. Using a biotynilated primer, each strand of the resulting PCR products carries biotin at its end (labeling). This labeled DNA strand can be separated and purified by covalent bonding to streptavidin. During the sequencing reaction, the four bases are added to the reaction solution separately. If incorporation occurs, an enzyme-reaction-based flash of light is emitted, which is detected in real time, similar to qPCR.
The crucial point is the quantification of nucleotide incorporation. This allows the degree of zygosity of a TSR position to be precisely determined.
Pyrosequencing is based on endpoint PCR. Using a biotynilated primer, each strand of the resulting PCR products carries biotin at its end (labeling). Through covalent binding to streptavidin, this labeled DNA strand can be separated and purified. During the sequencing reaction, the four bases are added to the reaction solution separately. If incorporation occurs, an enzyme-reaction-based flash of light is emitted, which, as in qPCR, is detected in real time.
The crucial point is the quantification of nucleotide incorporation. This allows the degree of zygosity of a TSR position to be precisely determined.
Endpoint PCR serves as the basis for fragment length analysis. By using a fluorophore-labeled primer, each strand of the resulting PCR products carries a fluorescent dye at its end (labeling).
High-resolution capillary electrophoresis, equipped with a detector unit for fluorescence signals, allows the lengths of the PCR fragments to be determined with nucleotide precision.
Fragment length analysis is used in the following analyses:
To identify methylated cytosines, the DNA must be subjected to a bisulfite reaction before endpoint PCR. In this reaction, a uracil is incorporated for all cytosines that are not methylated and are directly adjacent to a guanine base (CpG dinucleotide).
In the subsequent endpoint PCR, uracil is replaced by thymine, resulting in a different base sequence compared to the original.
The sequence can be determined after the end of the reaction either using the Sanger method or the pyro method.
Methylation analysis is used in the following applications:
In cooperation with two leading institutions, IDENTXX offers greenhouse biotests for determining metabolic resistance, also known as nTSR.
The partners have both a modern spraying facility and sufficient greenhouse capacity and qualified personnel.
The following points are particularly noteworthy:
contact
Address
information